wavepy.utils¶
Utility functions to help.
Functions:
crop_matrix_at_indexes(input_matrix, ...) |
Alias for np.copy(inputMatrix[i_min:i_max, j_min:j_max]) |
||
date_now_str() |
Returns the current date as a string in the format YYmmDD. | ||
datetime_now_str() |
Returns the current date and time as a string in the format YYmmDD_HHMMSS. | ||
dummy_images([imagetype, shape]) |
Dummy images for simple tests. | ||
find_nearest_value(input_array, value) |
Alias for | ||
find_nearest_value_index(input_array, value) |
Similar to wavepy.utils.find_nearest_value(), but returns |
||
fouriercoordmatrix(npointsx, deltax, ...) |
For back compability | ||
fouriercoordvec(npoints, delta) |
For back compability | ||
graphical_roi_idx(zmatrix[, verbose, ...]) |
Function to define a rectangular region of interest (ROI) in an image. | ||
graphical_select_point_idx(zmatrix[, ...]) |
Plot a 2D array and allow to pick a point in the image. | ||
h5_list_of_groups(h5file) |
Get the names of all groups and subgroups in a hdf5 file. | ||
load_ini_file(inifname) |
|
||
nan_mask_threshold(input_matrix[, threshold]) |
Calculate a square mask for array above OR below a threshold | ||
plot_profile(xmatrix, ymatrix, zmatrix[, ...]) |
Plot contourf in the main graph plus profiles over vertical and horizontal lines defined with mouse. | ||
print_blue(message) |
Print with colored characters. | ||
print_color(message[, color, highlights, attrs]) |
Print with colored characters. | ||
print_red(message) |
Print with colored characters. | ||
progress_bar4pmap(res[, sleep_time]) |
Progress bar from tqdm to be used with the function multiprocessing.starmap_async(). |
||
realcoordmatrix(npointsx, deltax, npointsy, ...) |
Build a matrix (2D array) with real space coordinates based on the number of points and bin (pixels) size. | ||
realcoordmatrix_fromvec(xvec, yvec) |
Alias for np.meshgrid(xvec, yvec) |
||
realcoordvec(npoints, delta) |
Build a vector with real space coordinates based on the number of points and bin (pixels) size. | ||
rotate_img_graphical(array2D[, order, mode, ...]) |
GUI to rotate an image | ||
select_dir([message_to_print, pattern]) |
List subdirectories of the current working directory, and expected the user to choose one of them. | ||
select_file([pattern, message_to_print]) |
List files under the subdirectories of the current working directory, and expected the user to choose one of them. | ||
time_now_str() |
Returns the current time as a string in the format HHMMSS. |
-
wavepy.utils.print_color(message, color='red', highlights='on_white', attrs='')[source]¶ Print with colored characters. It is only a alias for colored print using the package
termcolorand equals to:print(termcolor.colored(message, color, highlights, attrs=attrs))
See options at https://pypi.python.org/pypi/termcolor
Parameters: - message (str) – Message to print.
- color, highlights (str)
- attrs (list)
-
wavepy.utils.print_red(message)[source]¶ Print with colored characters. It is only a alias for colored print using the package
termcolorand equals to:print(termcolor.colored(message, color='red'))
Parameters: message (str) – Message to print.
-
wavepy.utils.print_blue(message)[source]¶ Print with colored characters. It is only a alias for colored print using the package
termcolorand equals to:print(termcolor.colored(message, color='blue'))
Parameters: message (str) – Message to print.
-
wavepy.utils.plot_profile(xmatrix, ymatrix, zmatrix, xlabel='x', ylabel='y', zlabel='z', title='Title', xo=None, yo=None, xunit='', yunit='', do_fwhm=True, arg4main=None, arg4top=None, arg4side=None)[source]¶ Plot contourf in the main graph plus profiles over vertical and horizontal lines defined with mouse.
Parameters: - xmatrix, ymatrix (ndarray) – x and y matrix coordinates generated with
numpy.meshgrid() - zmatrix (ndarray) – Matrix with the data. Note that
xmatrix,ymatrixandzmatrixmust have the same shape - xlabel, ylabel, zlabel (str, optional) – Labels for the axes
x,yandz. - title (str, optional) – title for the main graph #BUG: sometimes this title disappear
- xo, yo (float, optional) – if equal to
None, it allows to use the mouse to choose the vertical and horizontal lines for the profile. If notNone, the profiles lines are are centered at(xo,yo) - xunit, yunit (str, optional) – String to be shown after the values in the small text box
- do_fwhm (Boolean, optional) – Calculate and print the FWHM in the figure. The script to calculate the
FWHM is not very robust, it works well if only one well defined peak is
present. Turn this off by setting this var to
False - *arg4main – *args for the main graph
- *arg4top – *args for the top graph
- *arg4side – *args for the side graph
Returns: - ax_main, ax_top, ax_side (matplotlib.axes) – return the axes in case one wants to modify them.
- delta_x, delta_y (float)
Example
>>> import numpy as np >>> import wavepy.utils as wpu >>> xx, yy = np.meshgrid(np.linspace(-1, 1, 101), np.linspace(-1, 1, 101)) >>> wpu.plot_profile(xx, yy, np.exp(-(xx**2+yy**2)/.2))
Animation of the example above:
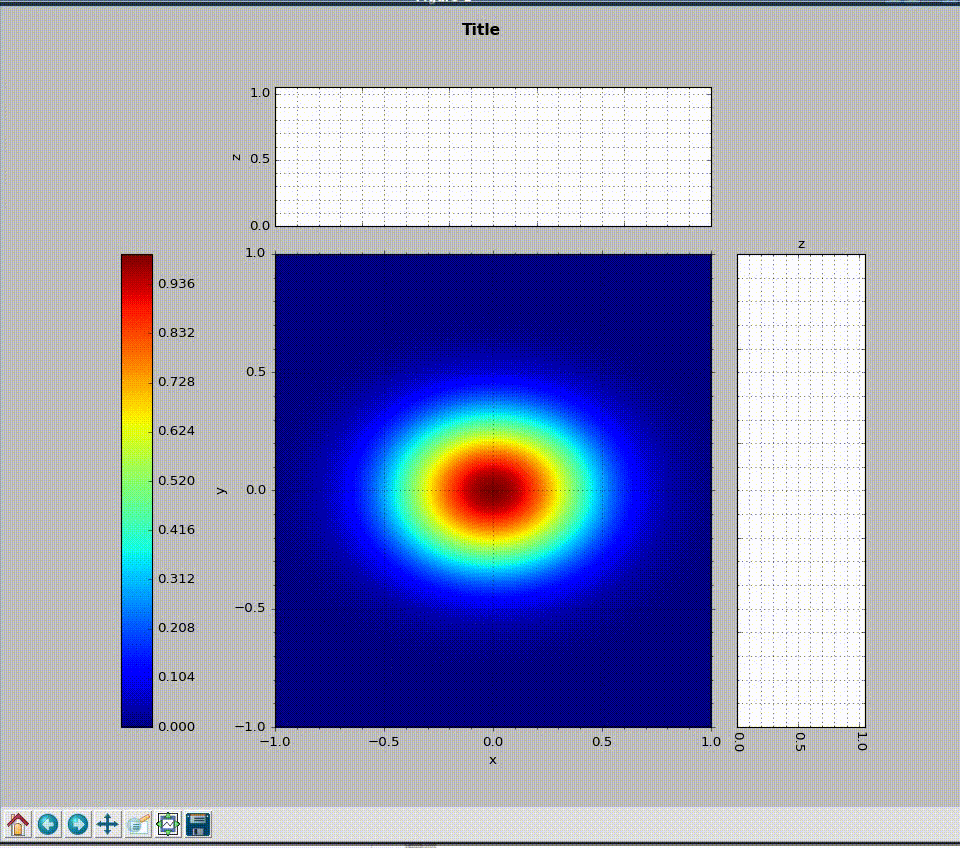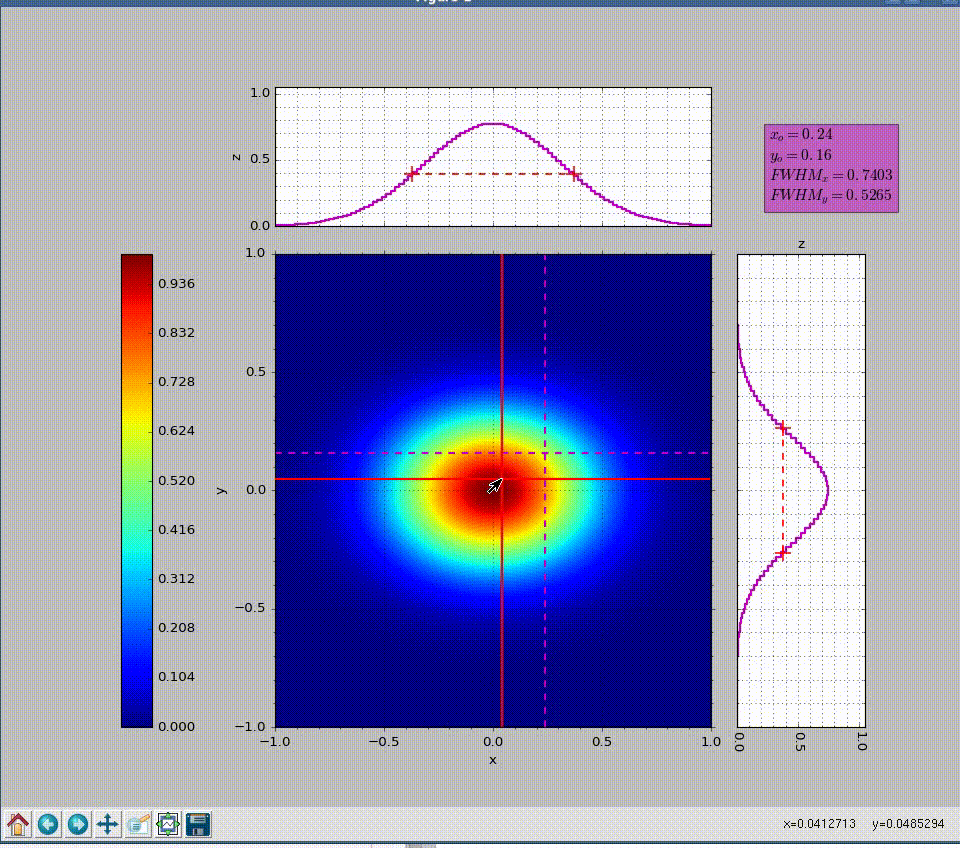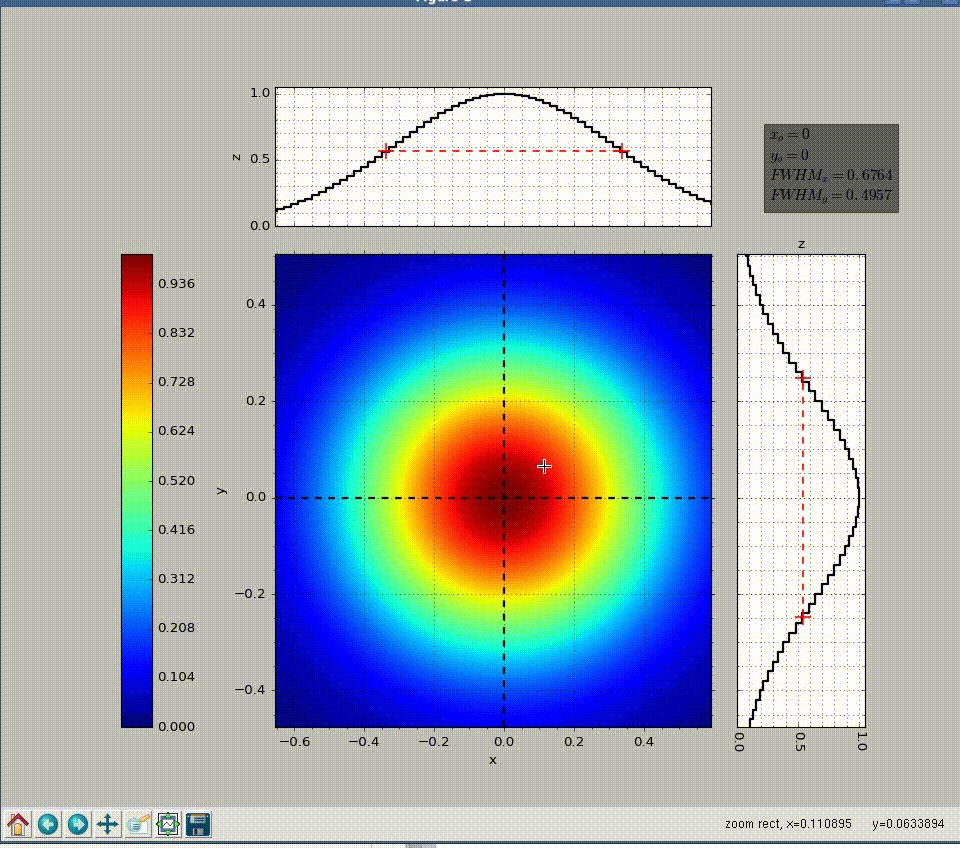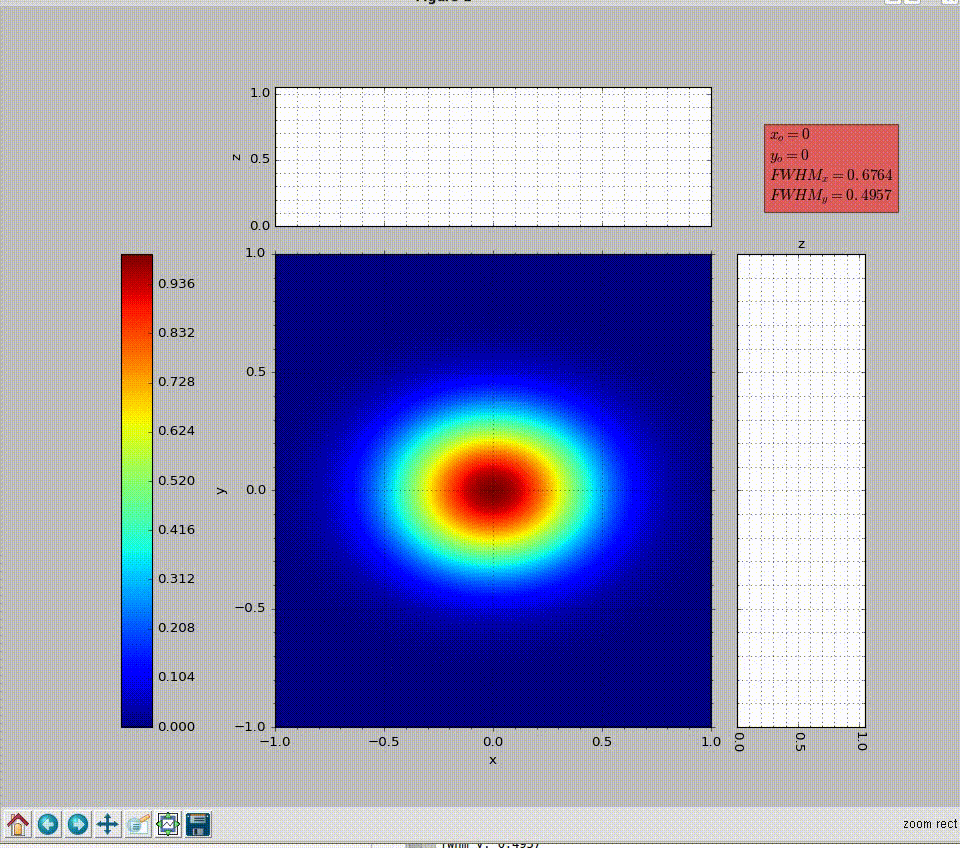
- xmatrix, ymatrix (ndarray) – x and y matrix coordinates generated with
-
wavepy.utils.select_file(pattern='*', message_to_print=None)[source]¶ List files under the subdirectories of the current working directory, and expected the user to choose one of them.
The list of files is of the form
number: filename. The user choose the file by typing the number of the desired filename.Parameters: - pattern (str) – list only files with this patter. Similar to pattern in the linux comands ls, grep, etc
- message_to_print (str, optional)
Returns: filename (str) – path and name of the file
Example
>>> select_file('*.dat')
-
wavepy.utils.select_dir(message_to_print=None, pattern='**/')[source]¶ List subdirectories of the current working directory, and expected the user to choose one of them.
The list of files is of the form
number: filename. The user choose the file by typing the number of the desired filename.Similar to
wavepy.utils.select_file()Parameters: message_to_print (str, optional) Returns: str – directory path See also
-
wavepy.utils.nan_mask_threshold(input_matrix, threshold=0.0)[source]¶ Calculate a square mask for array above OR below a threshold
Parameters: - input_matrix (ndarray) – 2 dimensional (or n-dimensional?) numpy.array to be masked
- threshold (float) – threshold for masking. If real (imaginary) value, values below(above) the threshold are set to NAN
Returns: ndarray – array with values either equal to 1 or NAN.
Example
To use as a mask for array use:
>>> mask = nan_mask_threshold(input_array, threshold) >>> masked_array = input_array*mask
Notes
- Note that
array[mask]will return only the values wheremask == 1. - Also note that this is NOT the same as the masked arrays in numpy.
-
wavepy.utils.crop_matrix_at_indexes(input_matrix, list_of_indexes)[source]¶ Alias for
np.copy(inputMatrix[i_min:i_max, j_min:j_max])Parameters: - input_matrix (ndarray) – 2 dimensional array
- list_of_indexes (list) – list in the format
[i_min, i_max, j_min, j_max]
Returns: ndarray – copy of the sub-region
inputMatrix[i_min:i_max, j_min:j_max]of the inputMatrix.Warning
Note the difference of copy and view in Numpy.
-
wavepy.utils.crop_graphic(xvec=None, yvec=None, zmatrix=None, verbose=False, kargs4graph={})[source]¶ Function to crop an image to the ROI selected using the mouse.
wavepy.utils.graphical_roi_idx()is first used to plot and select the ROI. The function then returns the croped version of the matrix, the cropped coordinate vectorsxandy, and the indexes[i_min, i_max, j_min,_j_max]Parameters: - xvec, yvec (1D ndarray) – vector with the coordinates
xandy. See below how the returned variables change dependnding whether these vectors are provided. - zmatrix (2D numpy array) – image to be croped, as an 2D ndarray
- **kargs4graph – kargs for main graph
Returns: - 1D ndarray, 1D ndarray – cropped coordinate vectors
xandy. These two vectors are only returned the input vectorsxvecandxvecare provided - 2D ndarray – cropped image
- list – indexes of the crop
[i_min, i_max, j_min,_j_max]. Useful when the same crop must be applies to other images
Examples
>>> import numpy as np >>> import matplotlib as plt >>> xVec = np.arange(0.,101) >>> yVec = np.arange(0.,101) >>> img = dummy_images('Shapes', size=(101,101), FWHM_x = .5, FWHM_y=1.0) >>> (imgCroped, idx4crop) = crop_graphic(zmatrix=img) >>> plt.imshow(imgCroped, cmap='Spectral') >>> (xVecCroped, >>> yVecCroped, >>> imgCroped, idx4crop) = crop_graphic(xVec, yVec, img) >>> plt.imshow(imgCroped, cmap='Spectral', >>> extent=np.array([xVecCroped[0], xVecCroped[-1], >>> yVecCroped[0], yVecCroped[-1]])
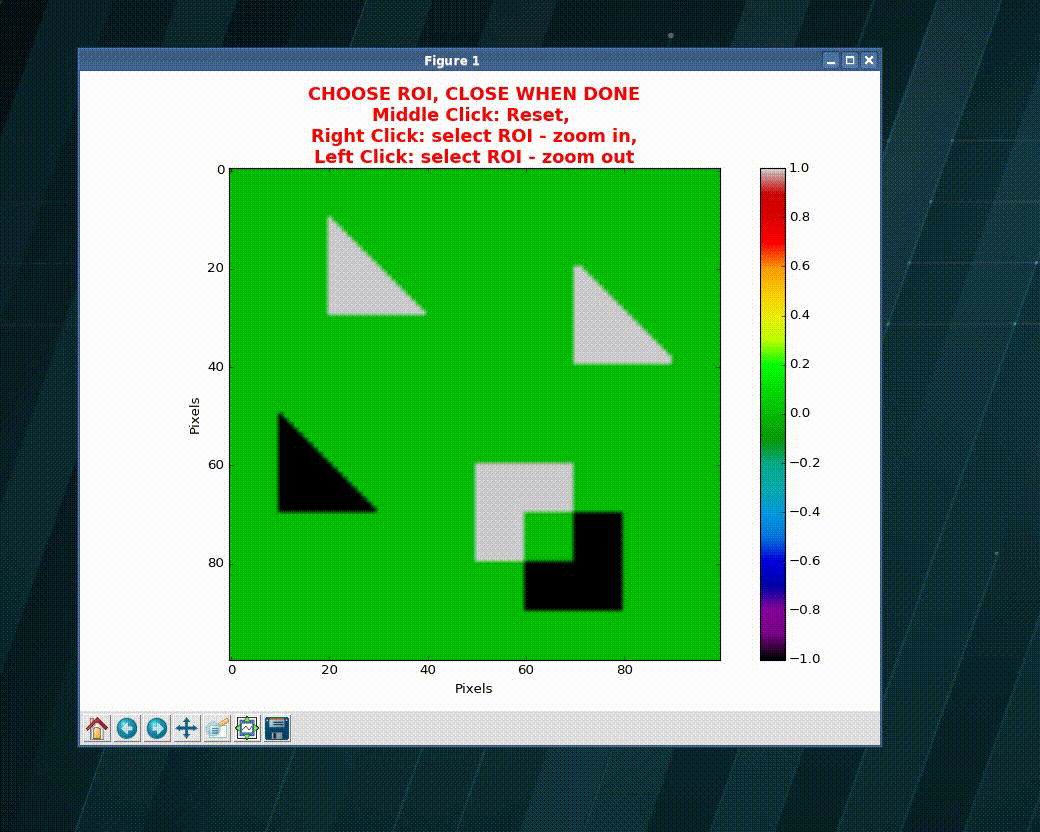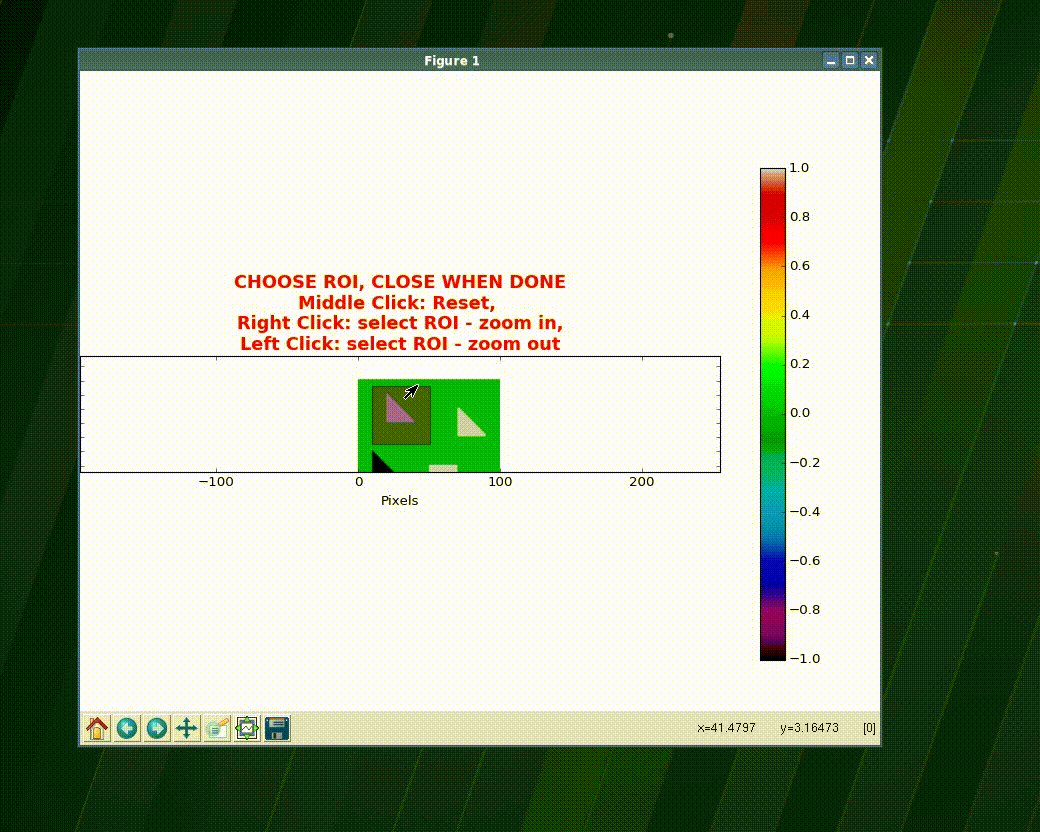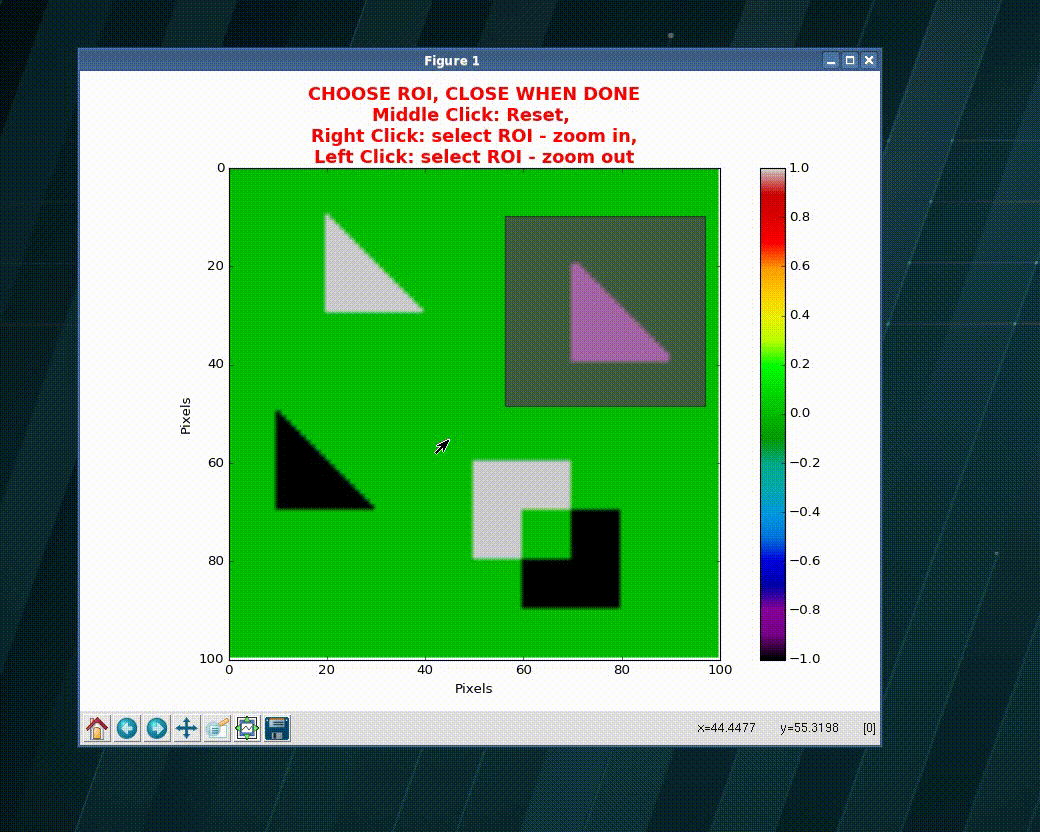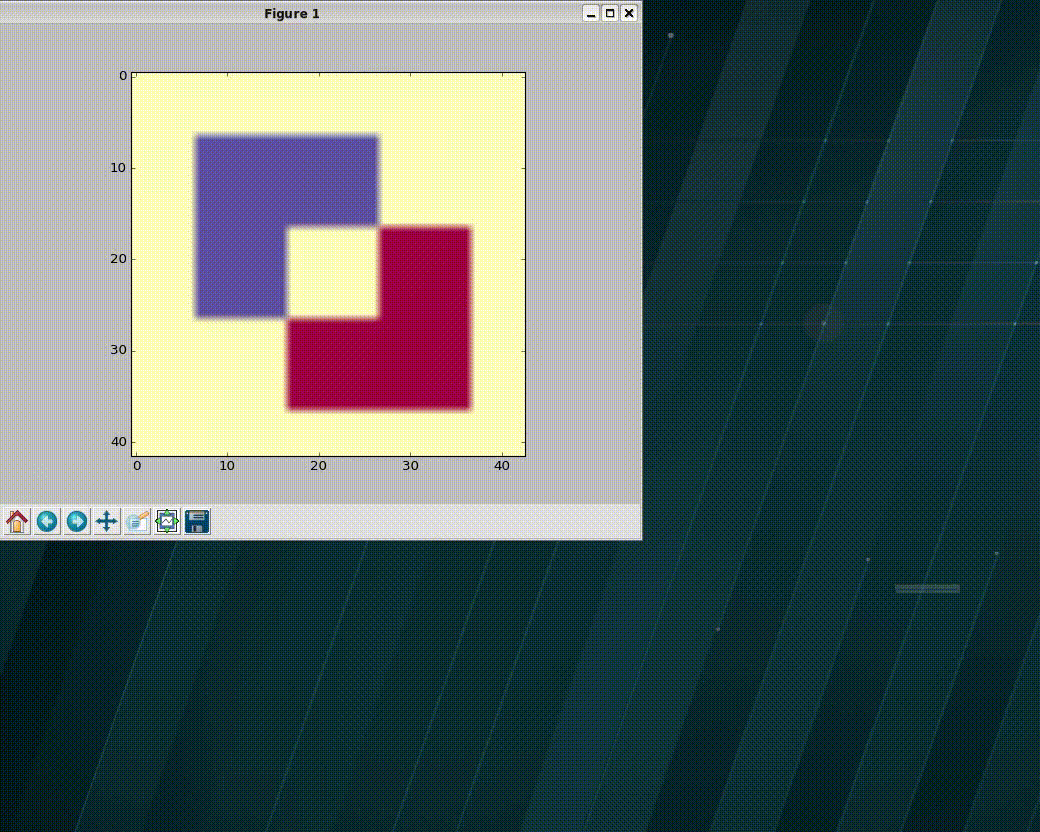
- xvec, yvec (1D ndarray) – vector with the coordinates
-
wavepy.utils.crop_graphic_image(image, verbose=False, **kargs4graph)[source]¶ Similar to
wavepy.utils.crop_graphic(), but only for the main matrix (and not for the x and y vectors). The function then returns the croped version of the image and the indexes[i_min, i_max, j_min,_j_max]Parameters: - zmatrix (2D numpy array) – image to be croped, as an 2D ndarray
- **kargs4graph – kargs for main graph
Returns: - 2D ndarray – cropped image
- list – indexes of the crop
[i_min, i_max, j_min,_j_max]. Useful when the same crop must be applies to other images
-
wavepy.utils.crop_graphic_image(image, verbose=False, **kargs4graph)[source] Similar to
wavepy.utils.crop_graphic(), but only for the main matrix (and not for the x and y vectors). The function then returns the croped version of the image and the indexes[i_min, i_max, j_min,_j_max]Parameters: - zmatrix (2D numpy array) – image to be croped, as an 2D ndarray
- **kargs4graph – kargs for main graph
Returns: - 2D ndarray – cropped image
- list – indexes of the crop
[i_min, i_max, j_min,_j_max]. Useful when the same crop must be applies to other images
-
wavepy.utils.graphical_select_point_idx(zmatrix, verbose=False, kargs4graph={})[source]¶ Plot a 2D array and allow to pick a point in the image. Returns the last selected position x and y of the choosen point
Parameters: - zmatrix (2D numpy array) – main image
- verbose (Boolean) – verbose mode
- **kargs4graph – kargs for main graph
Returns: int, int – two integers with the point indexes
xandyExample
>>> jo, io = graphical_select_point_idx(array2D) >>> value = array2D[io, jo]
See also
-
wavepy.utils.find_nearest_value(input_array, value)[source]¶ Alias for
input_array.flatten()[np.argmin(np.abs(input_array.flatten() - value))]In a array of float numbers, due to the precision, it is impossible to find exact values. For instance something like
array1[array2==0.0]might fail because the zero values in the float arrayarray2are actually something like 0.0004324235 (fictious value).This function will return the value in the array that is the nearest to the parameter
value.Parameters: - input_array (ndarray)
- value (float)
Returns: ndarray
Example
>>> foo = dummy_images('NormalDist') >>> find_nearest_value(foo, 0.5000) 0.50003537554879007
See also
wavepy:utils:find_nearest_value_index()
-
wavepy.utils.find_nearest_value_index(input_array, value)[source]¶ Similar to
wavepy.utils.find_nearest_value(), but returns the index of the nearest value (instead of the value itself)Parameters: - input_array (ndarray)
- value (float)
Returns: tuple of ndarray – each array have the index of the nearest value in each dimension
Note
In principle it has no limit of the number of dimensions.
Example
>>> foo = dummy_images('NormalDist') >>> find_nearest_value(foo, 0.5000) 0.50003537554879007 >>> (i_index, j_index) = find_nearest_value_index(foo, 0.500) >>> foo[i_index[:], j_index[:]] array([ 0.50003538, 0.50003538, 0.50003538, 0.50003538])
See also
wavepy:utils:find_nearest_value()
-
wavepy.utils.dummy_images(imagetype=None, shape=(100, 100), **kwargs)[source]¶ Dummy images for simple tests.
Parameters: - imagetype (str) – See options Below
- shape (tuple) – Shape of the image. Similar to
numpy.shape - kwargs – keyword arguments depending on the image type.
Image types
Noise (default): alias for
np.random.random(shape)Stripes:
kwargs: nLinesH, nLinesVSumOfHarmonics: image is defined by: .. math:: sum_{ij} Amp_{ij} cos (2 pi i y) cos (2 pi j x).
- Note that
xandyare assumed to in the range [-1, 1]. The keywordkwargs: harmAmplis a 2D list that can be used to set the values for Amp_ij, see Examples.
- Note that
Shapes: see Examples.
kwargs=noise, amplitude of noise to be added to the imageNormalDist: Normal distribution where it is assumed that
xandyare in the interval [-1,1].keywords: FWHM_x, FWHM_y
Returns: 2D ndarray Examples
>>> import matplotlib.pyplot as plt >>> plt.imshow(dummy_images())
is the same than
>>> plt.imshow(dummy_images('Noise'))
>>> plt.imshow(dummy_images('Stripes', nLinesV=5))

>>> plt.imshow(dummy_images('Stripes', nLinesH=8))

>>> plt.imshow(dummy_images('Checked', nLinesH=8, nLinesV=5))
>>> plt.imshow(dummy_images('SumOfHarmonics', harmAmpl=[[1,0,1],[0,1,0]]))
>>> plt.imshow(dummy_images('Shapes', noise = 1))
>>> plt.imshow(dummy_images('NormalDist', FWHM_x = .5, FWHM_y=1.0))
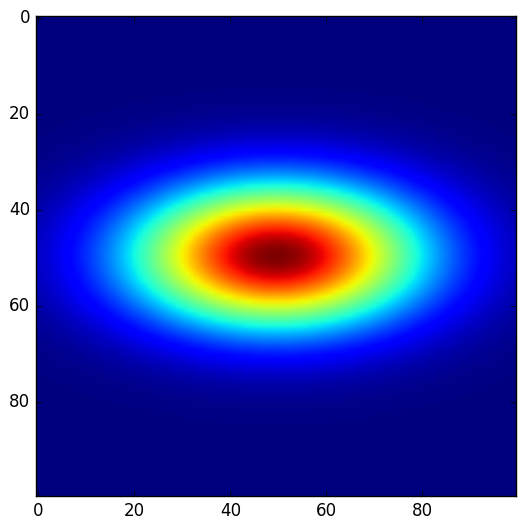
-
wavepy.utils.graphical_roi_idx(zmatrix, verbose=False, kargs4graph={})[source]¶ Function to define a rectangular region of interest (ROI) in an image.
The image is plotted and, using the mouse, the user select the region of interest (ROI). The ROI is ploted as an transparent rectangular region. When the image is closed the function returns the indexes
[i_min, i_max, j_min, j_max]of the ROI.Parameters: - input_array (ndarray)
- verbose (Boolean) – In the verbose mode it is printed some additional infomations, like the ROI indexes, as the user select different ROI’s
- **kargs4graph – Options for the main graph. WARNING: not tested very well
Returns: list – indexes of the ROI
[i_min, i_max, j_min,_j_max]. Useful when the same crop must be applies to other imagesNote
In principle it has no limit of the number of dimensions.
Example
See example at
wavepy:utils:crop_graphic()See also
wavepy:utils:crop_graphic()
-
wavepy.utils.crop_graphic(xvec=None, yvec=None, zmatrix=None, verbose=False, kargs4graph={})[source] Function to crop an image to the ROI selected using the mouse.
wavepy.utils.graphical_roi_idx()is first used to plot and select the ROI. The function then returns the croped version of the matrix, the cropped coordinate vectorsxandy, and the indexes[i_min, i_max, j_min,_j_max]Parameters: - xvec, yvec (1D ndarray) – vector with the coordinates
xandy. See below how the returned variables change dependnding whether these vectors are provided. - zmatrix (2D numpy array) – image to be croped, as an 2D ndarray
- **kargs4graph – kargs for main graph
Returns: - 1D ndarray, 1D ndarray – cropped coordinate vectors
xandy. These two vectors are only returned the input vectorsxvecandxvecare provided - 2D ndarray – cropped image
- list – indexes of the crop
[i_min, i_max, j_min,_j_max]. Useful when the same crop must be applies to other images
Examples
>>> import numpy as np >>> import matplotlib as plt >>> xVec = np.arange(0.,101) >>> yVec = np.arange(0.,101) >>> img = dummy_images('Shapes', size=(101,101), FWHM_x = .5, FWHM_y=1.0) >>> (imgCroped, idx4crop) = crop_graphic(zmatrix=img) >>> plt.imshow(imgCroped, cmap='Spectral') >>> (xVecCroped, >>> yVecCroped, >>> imgCroped, idx4crop) = crop_graphic(xVec, yVec, img) >>> plt.imshow(imgCroped, cmap='Spectral', >>> extent=np.array([xVecCroped[0], xVecCroped[-1], >>> yVecCroped[0], yVecCroped[-1]])

- xvec, yvec (1D ndarray) – vector with the coordinates
-
wavepy.utils.choose_unit(array)[source]¶ Script to choose good(best) units in engineering notation for a
ndarray.For a given input array, the function returns
factorandunitaccording to\[10^{n} < \max(array) < 10^{n + 3}\]n factor (float) unit(str) 0 1.0 ''empty string-9 10^-9 n-6 10^-6 r'\mu'-3 10^-3 m+3 10^-6 k+6 10^-9 M+9 10^-6 Gn=-6returns\musince this is the latex syntax for micro. See Example.Parameters: array (ndarray) – array from where to choose proper unit. Returns: float, unit – Multiplication Factor and strig for unit Example
>>> array1 = np.linspace(0,100e-6,101) >>> array2 = array1*1e10 >>> factor1, unit1 = choose_unit(array1) >>> factor2, unit2 = choose_unit(array2) >>> plt.plot(array1*factor1,array2*factor2) >>> plt.xlabel(r'${0} m$'.format(unit1)) >>> plt.ylabel(r'${0} m$'.format(unit2))
The syntax
r'$ string $ 'is necessary to use latex commands in thematplotliblabels.
-
wavepy.utils.datetime_now_str()[source]¶ Returns the current date and time as a string in the format YYmmDD_HHMMSS. Alias for
time.strftime("%Y%m%d_%H%M%S").Returns: str
-
wavepy.utils.time_now_str()[source]¶ Returns the current time as a string in the format HHMMSS. Alias for
time.strftime("%H%M%S").Returns: str
-
wavepy.utils.date_now_str()[source]¶ Returns the current date as a string in the format YYmmDD. Alias for
time.strftime("%Y%m%d").Returns: str
-
wavepy.utils.realcoordvec(npoints, delta)[source]¶ Build a vector with real space coordinates based on the number of points and bin (pixels) size.
Alias for
np.mgrid[-npoints/2*delta:npoints/2*delta-delta:npoints*1j]Parameters: - npoints (int) – number of points
- delta (float) – vector with the values of x
Returns: ndarray – vector (1D array) with real coordinates
Example
>>> realcoordvec(10,1) array([-5., -4., -3., -2., -1., 0., 1., 2., 3., 4.])
-
wavepy.utils.realcoordmatrix_fromvec(xvec, yvec)[source]¶ Alias for
np.meshgrid(xvec, yvec)Parameters: xvec, yvec (ndarray) – vector (1D array) with real coordinates Returns: ndarray – 2 matrices (1D array) with real coordinates Example
>>> vecx = realcoordvec(3,1) >>> vecy = realcoordvec(4,1) >>> realcoordmatrix_fromvec(vecx, vecy) [array([[-1.5, -0.5, 0.5], [-1.5, -0.5, 0.5], [-1.5, -0.5, 0.5], [-1.5, -0.5, 0.5]]), array([[-2., -2., -2.], [-1., -1., -1.], [ 0., 0., 0.], [ 1., 1., 1.]])]
-
wavepy.utils.realcoordmatrix(npointsx, deltax, npointsy, deltay)[source]¶ Build a matrix (2D array) with real space coordinates based on the number of points and bin (pixels) size.
Alias for
realcoordmatrix_fromvec(realcoordvec(nx, delx), realcoordvec(ny, dely))Parameters: - npointsx, npointsy (int) – number of points in the x and y directions
- deltax, deltay (float) – step size in the x and y directions
Returns: ndarray, ndarray – 2 matrices (2D array) with real coordinates
Example
>>> realcoordmatrix(3,1,4,1) [array([[-1.5, -0.5, 0.5], [-1.5, -0.5, 0.5], [-1.5, -0.5, 0.5], [-1.5, -0.5, 0.5]]), array([[-2., -2., -2.], [-1., -1., -1.], [ 0., 0., 0.], [ 1., 1., 1.]])]
-
wavepy.utils.reciprocalcoordvec(npoints, delta)[source]¶ Create coordinates in the (spatial) frequency domain based on the number of points
nand the step (binning)\Delta xin the REAL SPACE. It returns a vector of frequencies with values in the interval\[\left[ \frac{-1}{2 \Delta x} : \frac{1}{2 \Delta x} - \frac{1}{n \Delta x} \right],\]with the same number of points.
Parameters: - npoints (int) – number of points
- delta (float) – step size in the REAL SPACE
Returns: ndarray
Example
>>> reciprocalcoordvec(10,1e-3) array([-500., -400., -300., -200., -100., 0., 100., 200., 300., 400.])
-
wavepy.utils.reciprocalcoordmatrix(npointsx, deltax, npointsy, deltay)[source]¶ Similar to
wavepy.utils.reciprocalcoordvec(), but for matrices (2D arrays).Parameters: - npointsx, npointsy (int) – number of points in the x and y directions
- deltax, deltay (float) – step size in the x and y directions
Returns: ndarray, ndarray – 2 matrices (2D array) with coordinates in the frequencies domain
Example
>>> reciprocalcoordmatrix(5, 1e-3, 4, 1e-3) [array([[-500., -300., -100., 100., 300.], [-500., -300., -100., 100., 300.], [-500., -300., -100., 100., 300.], [-500., -300., -100., 100., 300.]]), array([[-500., -500., -500., -500., -500.], [-250., -250., -250., -250., -250.], [ 0., 0., 0., 0., 0.], [ 250., 250., 250., 250., 250.]])]
-
wavepy.utils.h5_list_of_groups(h5file)[source]¶ Get the names of all groups and subgroups in a hdf5 file.
Parameters: h5file (h5py file) Returns: list – list of strings with group names Example
>>> fh5 = h5py.File(filename,'r') >>> listOfGoups = h5_list_of_groups(fh5) >>> for group in listOfGoups: print(group)
-
wavepy.utils.progress_bar4pmap(res, sleep_time=1.0)[source]¶ Progress bar from
tqdmto be used with the functionmultiprocessing.starmap_async().It holds the program in a loop waiting
multiprocessing.starmap_async()to finishParameters: - res (result object of the
multiprocessing.Poolclass) - sleep_time
Example
>>> from multiprocessing import Pool >>> p = Pool() >>> res = p.starmap_async(...) # use your function inside brackets >>> p.close() # No more work >>> progress_bar4pmap(res)
- res (result object of the
-
wavepy.utils.load_ini_file(inifname)[source]¶ Parameters: inifname (str) – name of the *.inifile.Returns: configparser objects Example
Example of
inifile:[Files] image_filename = file1.tif ref_filename = file2.tif [Parameters] par1 = 10.5e-5 par2 = 10, 100, 500, 600 par can have long name = 25 par3 = the value can be anything
If we create a file named
.temp.iniwith the example above, we can load it as:>>> config = load_ini_file('.temp.ini') >>> print(config['Parameters']['Par1'] )
See also
wavepy.utils.load_ini_file_terminal_dialog(),wavepy.utils.get_from_ini_file(),wavepy.utils.set_at_ini_file().
-
wavepy.utils.rocking_3d_figure(ax, outfname='out.ogv', elevAmp=50, azimAmpl=90, elevOffset=0, azimOffset=0, dpi=50, npoints=200, del_tmp_imgs=True)[source]¶ Saves an image at different view angles and join the images to make a short animation. The movement is defined by setting the elevation and azimut angles following a sine function. The frequency of the vertical movement (elevation) is twice of the horizontal (azimute), forming a figure eight movement.
Parameters: - ax (3D axis object) – See example below how to create this object. If None, this will use the temporary images from a previous run
- outfname (str) – output file name. Note that the extension defines the file format. This function has been tested for the formats .gif (not recomended, big files and poor quality), .mp4, .mkv and .ogv. For LibreOffice, .ogv is the recomend format.
- elevAmp (float) – amplitude of elevation movement, in degrees
- azimAmpl (float) – amplitude of azimutal movement, in degrees. If negative, the image will continually rotate around the azimute direction (no azimute rocking)
- elevOffset (float) – offset of elevation movement, in degrees
- azimOffset (float) – offset of azimutal movement, in degrees
- dpi (float) – resolution of the individual images
- npoints (int) – number of intermediary images to form the animation. More images will make the the movement slower and the animation longer.
- remove_images (float) – the program creates npoints temporary images, and this flag defines if these images are deleted or not
Example
>>> fig = plt.figure() >>> ax = fig.add_subplot(111, projection='3d') >>> xx = np.random.rand(npoints)*2-1 >>> yy = np.random.rand(npoints)*2-1 >>> zz = np.sinc(-(xx**2/.5**2+yy**2/1**2)) >>> ax.plot_trisurf(xx, yy, zz, cmap='viridis', linewidth=0.2, alpha = 0.8) >>> plt.show() >>> plt.pause(.5) >>> # this pause is necessary to plot the first image in the main screen >>> rocking_3d_figure(ax, 'out_080.ogv', elevAmp=45, azimAmpl=45, elevOffset=0, azimOffset=45, dpi=80) >>> # example of use of the del_tmp_imgs flag >>> rocking_3d_figure(ax, 'out_050.ogv', elevAmp=60, azimAmpl=60, elevOffset=10, azimOffset=45, dpi=50, del_tmp_imgs=False) >>> rocking_3d_figure(None, 'out2_050.gif', del_tmp_imgs=True)